What is ARIA?
I realise I've been working on it for years, but never really talked about it. So, what is this ARIA thing anyway? I hear you ask...

I realise that I don’t think I’ve really talked about what I do in my day job at all on this platform, and this is probably remiss of me, as it is actually quite exciting.
So, for the past… number of years… I’ve been working, among other things, as Head of IT at Instruct, which is the pan-European research infrastructure for structural biology. I direct the Instruct Digital Services team operating out of the Instruct Hub, and lead development on among other things, the ARIA platform which is our flagship Access and research data management platform. It is interesting, impactful and variable work where I work with some of the leading scientists in the field, and which lets me scratch my wanderlust!
So, what is ARIA then? Well, ARIA is a cloud platform we’ve developed at Instruct that does Access (and increasingly data management) for Research Infrastructures (RIs) and European science projects. We developed it out of a need at Instruct to manage researchers visiting our various research facilities in our member countries, manage the scientific proposal, conduct scientific and technical evaluation and collect feedback, and for the facilities themselves to be able to organise the visit at their end. It solved a real need within Instruct and streamlines our operations.
As we’ve grown we’ve been found that others have similar problems… as much as people might think they’re beautiful unique snowflakes, Access is basically Access, and so we offer ARIA to RIs and projects for a fee to cover the costs of our operation.
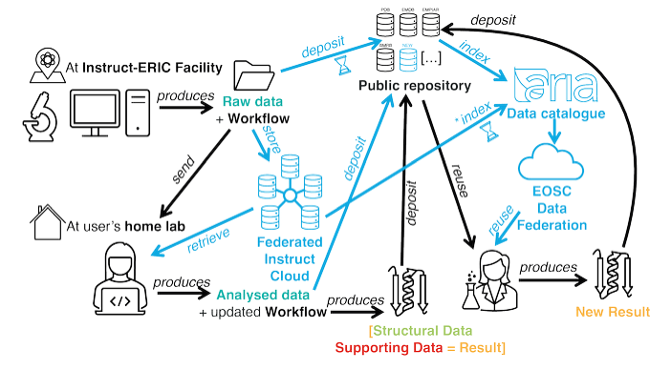
Increasingly, as part of our lead role in the Fragment Screen project, we have been expanding into data management in order to build out the wider Instruct data management plan. The issue always has been that once a researcher conducts their science at one of our facilities, they are often handed their data on a pen drive, at which point we lose oversight. As an organisation we have a commitment to FAIR open science, and so this presents a problem. So, using a tool called FandanGO, which works together with the ARIA platform, we are increasingly capturing experimental metadata, sample information, machine settings etc, together with the raw data location and any outputs (PDB depositions etc), and then associating it with the original scientific proposal. To help our smaller facilities, we are also piloting storage for raw data provision as part of this ecosystem, here FandanGO will talk to ARIA for a number of storage providers in our “federated cloud”.
This is pivotal as it gives us a complete cradle to grave overview of the research being done, and also allows us to cross reference it with other research on the same proteins. All this data can be used to feed AI models, all leading to faster drug discovery in the future, and will eventually be exposed by our connections to the wider European open science community via our connections with EOSC (which I am also working on, but that’s a whole other story). We are also working with international partners to link all of this up to other international data sharing networks out of the US and beyond.
Exciting stuff!
Anyway, I’ll write more about this going forward if folks are interested. Also, if you happen to be running an ERIC, research infrastructure, or are writing a project grant, in the life sciences or not, and you need Access or data management, then please reach out!
Leave a Reply